1) Install IntelliJ using Ubuntu Software Center This is the most recommended way of installing the IntelliJ IDE on Ubuntu. It is similar in functionality to other molecule drawing programs such as ChemDraw (TM, CambridgeSoft). Option 1 Installing Node.js with Apt from the Default Repositories Ubuntu 20.04 contains a version of Node.js in its default repositories that can be used to provide a consistent experience across multiple systems. Run terminal and then first, issue an update command. Before you install Docker Engine for the first time on a new host machine, you need to set up the Docker repository. Tutorial exercise: Cinnamaldehyde It is still easy to install. The use of these tools is summarized in the Toolbar reference. The first thing to do is log in to your Ubuntu instance and add the necessary repository (as the version of Docker found in the standard. 
Tutorial Install And Configure OpenVAS On Ubuntu 20.04. Initialize Cloud SDK After you install Cloud SDK successfully on your system. Follow the installation progress on the terminal screen. Step 1: Install Kubernetes In this step, we will be installing Kubernetes. sktop - 1:1.11.0-3 amd64 armhf arm64 ppc64el riscv64 s390x - Type: desktop-application ID: sktop Package: xdrawchem Name: C: xdrawchem Summary: fr_FR: diteur de structures chimiques et de ractions C: Chemical structures and reactions editor Description: C: >- Xdrawchem is a 2D editor for. This has several disadvantages, like poor markup, too technical descriptions for users of software centers, different components having the same description, etc. It operates in much the same fashion, and should be highly familiar to users of ChemDraw. Create a new Rancher server container using the following command: sudo docker run -d -restart=unless-stopped -p 8080:8080 rancher/server:stable. On Ubuntu and Debian systems, the Apache package and the service is called apache2. BLAST can be run standalone, as an NCBI Web client, via a local BLAST server and via a Web form at the NCBI Website.But fret not! Install Xfce desktop on Ubuntu using xfce4 package That's really simple. BLAST can be used to infer functional and evolutionary relationships between sequences as well as help identify members of gene families. The program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches. 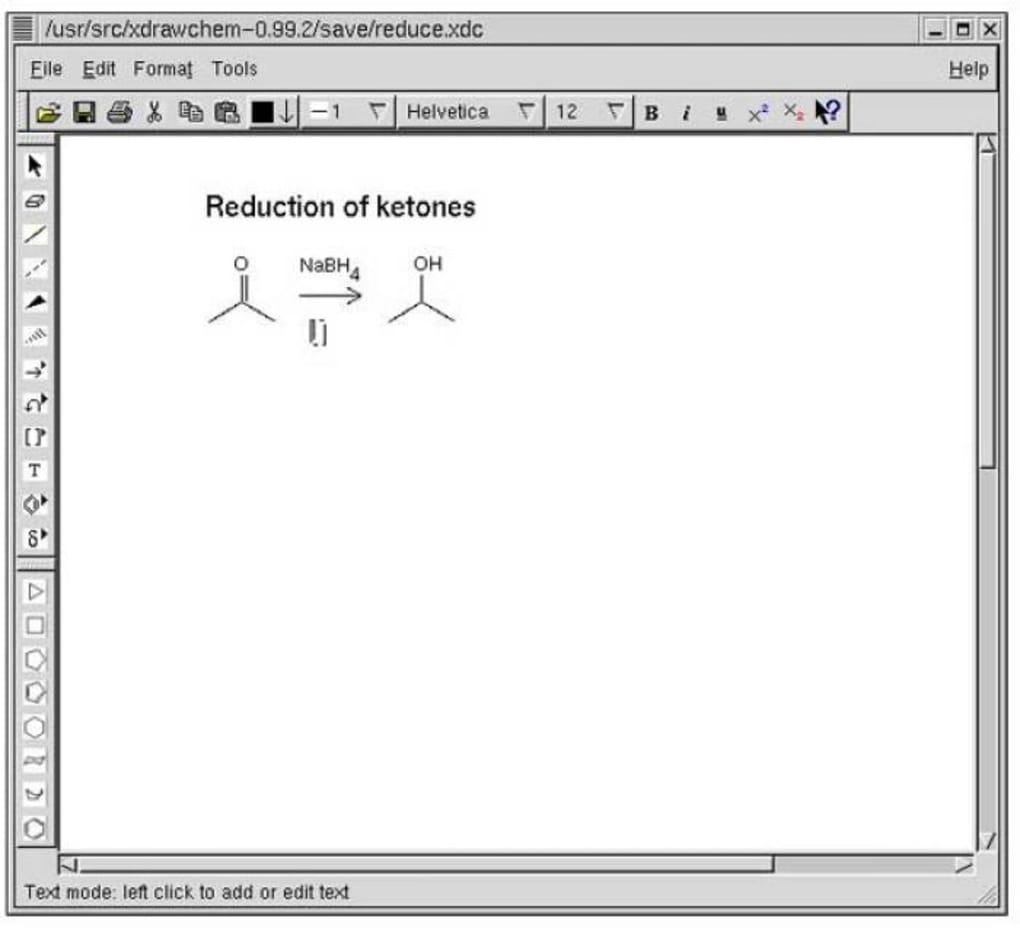
See also: BLAST, CLC Sequence Viewer, clustalw, COPASI, Cytoscape, emboss, Geneious, GENSCAN, hmmer, Integrative Genomics Viewer, meme/mast, Osprey, phylip, primer3, python, R, SnapGene, UGENE | BLASTĭescription: BLAST (Basic Local Alignment Search Tool) is an application for finding regions of local similarity between sequences. To run as a stand-alone application, you can write python scripts that contain Biopythoncommands See also: clustalw, GENSCAN, NetPBM Utilities, xv | Biopythonĭescription: collection of Python tools for computational molecular biology- allows processing of biological sequences and databasesįollowed by commands from the Biopython package, for example: Local documentation is locate at /mit/seven/doc/AFNI98 there is also a central information site and a resource site Please see documentation for usage of utility commands Supplied with many associated command-line utility functionsĪthena% *afni *_data-directory _(main application)Īthena% command options _files _(utilities) 
What Runs Where on Athena - Simulation and Modelingĭescription: collection of programs for viewing and analyzing functional magnetic resonance images of the brain. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
